|
2/4/2024 0 Comments Svm vs random forest
Boosting iteratively adds basis functions in a greedy fashion such that each additional basis function further reduces the selected loss (error) function : Gradient-boosted models can also handle interactions, automatically select variables, are robust to outliers, missing data and numerous correlated and irrelevant variables and can construct variable importance in exactly the same way as RF. Stochastic Gradient Boostingīoosting is an ensemble learning method for improving the predictive performance of classification or regression procedures, such as decision trees. We ranked SNPs by the relative importance of their contributions to predictive accuracy, quantified by how much prediction error increased when the observations left out of the bootstrap samples, the out-of-bag data for a SNP, were randomly permuted while data for all the other SNPs were left unchanged. 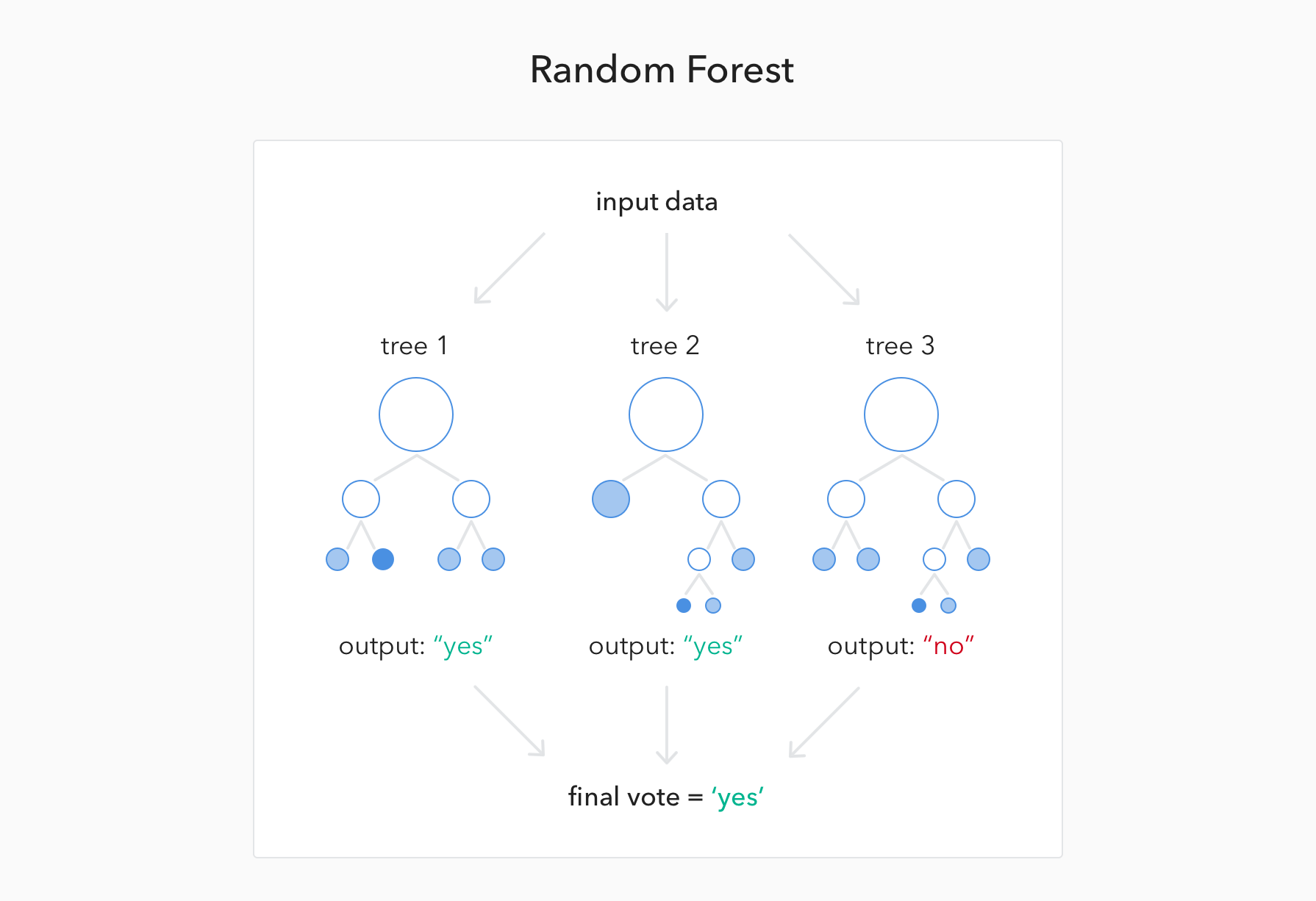
The parameter configuration with the highest prediction accuracy was ntree =1000, mtry = 3000 and nodesize =1. Following various recommendations, we evaluated different combinations of the values of the number of trees to grow, ntree = × the default value of mtry of sample size/3 for regression, and the minimum size of terminal nodes of trees, below which no split is attempted, nodesize = 1. We implemented RF in the R package randomForest with decision trees as base learners. Where Ψ b characterizes the bth RF tree in terms of split variables, cutpoints at each node, and terminal node values. RF regression was used to rank the SNPs in terms of their predictive importance. We comparatively evaluated the predictive performance of the three machine learning methods and RR-BLUP for estimating GEBVs using the common dataset simulated for the QTLMAS 2010 workshop. SVMs perform robustified regression using kernel functions of inner products of predictors. 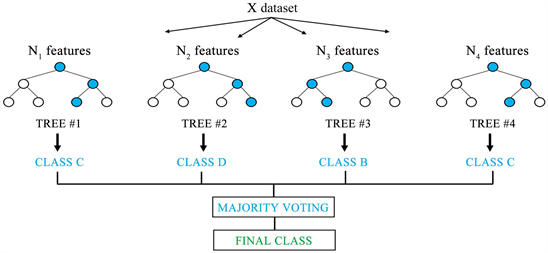
Boosting is a stagewise additive model fitting procedure that can enhance the predictive performance of weak learning algorithms. RF has several appealing properties that make it potentially attractive for GS : ( i) the number of markers can far exceed that of observations, ( ii) all markers, including those with weak effects, highly correlated and interacting markers have a chance to contribute to the model fit, ( iii) complex interactions between markers can be easily accommodated, ( iv) they can perform both simple and complex classification and regression accurately, ( v) they often require modest fine-tuning of parameters and the default parameterization often performs well, and ( vi) they make no distributional assumptions about the predictor variables. Here, we compare predictive performances among three of the most powerful machine learning methods with demonstrated high predictive accuracies in many application domains, namely RF boosting and SVMs and with RR-BLUP for predicting breeding values for quantitative traits. Given the wide range of approaches for predicting GEBVs, it is important to evaluate their performance, pros and cons to identify those able to accurately predict GEBVs. Genomic selection is a method for estimating GEBVs using dense molecular markers spanning the entire genome. ConclusionsĪccuracy was highest for boosting, intermediate for SVMs and lowest for RF but differed little among the three methods and relative to ridge regression BLUP (RR-BLUP). The correlations between the predicted and true breeding values were 0.547 for boosting, 0.497 for SVMs, and 0.483 for RF, indicating better performance for boosting than for SVMs and RF. The importance of each marker was ranked using RF and plotted against the position of the marker and associated QTLs on one of five simulated chromosomes. Predictive accuracy was measured as the Pearson correlation between GEBVs and observed values using 5-fold cross-validation and between predicted and true breeding values. We predicted GEBVs for one quantitative trait in a dataset simulated for the QTLMAS 2010 workshop. We evaluated the predictive accuracy of random forests (RF), stochastic gradient boosting (boosting) and support vector machines (SVMs) for predicting genomic breeding values using dense SNP markers and explored the utility of RF for ranking the predictive importance of markers for pre-screening markers or discovering chromosomal locations of QTLs. The existence of a wide array of marker-based approaches for predicting breeding values makes it essential to evaluate and compare their relative predictive performances to identify approaches able to accurately predict breeding values. Accurate prediction of genomic breeding values (GEBVs) presents a central challenge to contemporary plant and animal breeders. Genomic selection (GS) involves estimating breeding values using molecular markers spanning the entire genome.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
